Intriguing Study Reveals Genes And Languages Aren’t Always Found Together – What Does This Say About Our Ancestors?
Jan Bartek - AncientPages.com - More than 7,000 languages are spoken in the world. This linguistic diversity is passed on from one generation to the next, similarly to biological traits. But have language and genes evolved in parallel over the past few thousand years, as Charles Darwin originally thought?
An interdisciplinary team at the University of Zurich, together with the Max Planck Institute for Evolutionary Anthropology in Leipzig (Germany) has now examined this question at a global level. The researchers have developed a global database linking linguistic and genetic data entitled GeLaTo (Genes and Languages Together), which contains genetic information from some 4,000 individuals speaking 295 languages and representing 397 genetic populations.
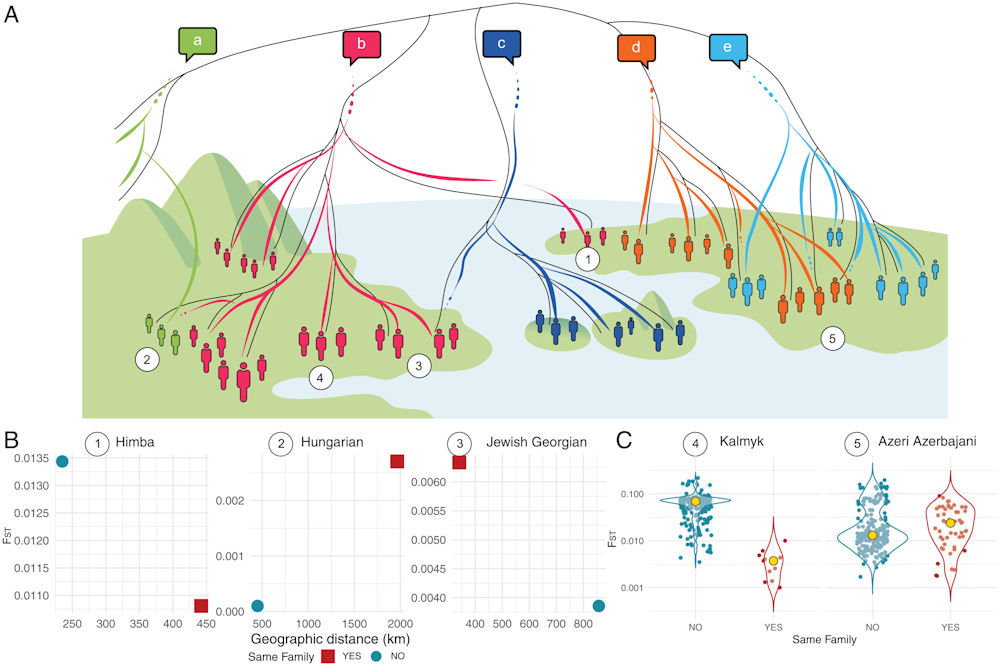
Overview of linguistic and genetic similarity. (A) Schematic illustration of possible scenarios of matches and mismatches in the transmission of genes and linguistic traits. Genetic (demographic) history is represented by solid black lines that differentiate groups of people (represented by human shapes). Linguistic history is represented by colored lines, differentiating five language families (a–e). The linguistic histories sometimes move in parallel with the demographic history and sometimes not. Numbers correspond to the different cases distinguished in B and C: 1. linguistic and genetic (matching) enclave; 2. linguistic mismatch (linguistic enclave); 3. genetic mismatch (genetic enclave); 4. population with genetic distances aligned with their linguistic relatives (matching profile); 5. population with genetic distances misaligned to their linguistic relatives (mismatching profile). (B) Examples of a heuristic associated with the three enclave cases shown in A. For each target population, we display the two smallest FST distances, respectively, to a population from the same family and a population from a different language family, together with their geographic distance. Himba (Atlantic-Congo family) fulfills the criteria of a matching enclave; Hungarian (Uralic family) fulfills the criteria of a linguistic enclave; Jewish Georgian (Kartvelian family) fulfills the criteria of a genetic enclave. (C) Examples of aligned and misaligned cases shown in A. For each population, the FST distribution within speakers of the same language family is compared with the FST distribution between the speakers of other language families. The yellow dot indicates the median. Kalmyk (Mongolic-Khitan) is aligned (i.e., is genetically closer) to speakers of the same language family; Azeri Azerbaijani (Turkic family) is misaligned to speakers of the same language family. FST distances are displayed on a logarithmically transformed scale. Credit: Proceedings of the National Academy of Sciences (2022). DOI: 10.1073/pnas.2122084119
One in five gene-language links point to language shifts
In their study, the researchers examined the extent to which the linguistic and genetic histories of populations coincided. People who speak related languages tend to also be genetically related, but this isn't always the case. "We focused on cases where the biological and linguistic patterns differed and investigated how often and where these mismatches occur," says Chiara Barbieri, UZH geneticist who led the study and initiated it together with colleagues when she was a postdoc at the Max-Planck-Institute.
The researchers found that about every fifth gene-language relation is a mismatch, and they occur worldwide. These mismatches can provide insights into the history of human evolution. "Once we know where such language shifts happened, we can better reconstruct how languages and populations spread across the world," says Balthasar Bickel, director of the National Center of Competence in Research (NCCR) Evolving Language, who co-supervised the study.
Switching to the local lingo
Most mismatches result from populations shifting to the language of a neighboring population that is genetically different. Some peoples on the tropical eastern slopes of the Andes speak a Quechua idiom that is typically spoken by groups with a different genetic profile who live at higher altitudes. The Damara people in Namibia, who are genetically related to the Bantu, communicate using a Khoe language that is spoken by genetically distant groups in the same area. And some hunter-gatherers who live in Central Africa speak predominantly Bantu languages without a strong genetic relatedness to the neighboring Bantu populations.
In addition, there are cases where migrants have picked up the local language of their new homes. The Jewish population in Georgia, for example, has adopted a South Caucasian language, while the Cochin Jews in India speak a Dravidian language. The case of Malta reflects its history as an island between two continents: While the Maltese are closely related to the people of Sicily, they speak an Afroasiatic language that is influenced by various Turkish and Indo-European languages.
Preserving their linguistic identity
"It appears that giving up your language isn't that difficult, also for practical reasons," says the last author Kentaro Shimizu, director of the URPP Evolution in Action: From Genomes to Ecosystems. However, it's more rare for people to preserve their original linguistic identity despite genetic assimilation with their neighbors. "Hungarian people, for example, are genetically similar to their neighbors, but their language is related to languages spoken in Siberia," Shintaro notes.
This makes Hungarian speakers stand out from among the rest of Europe and parts of Asia, where most people speak Indo-European languages, such as French, German, Hindi, Farsi, Greek and many others. Indo-European has not only been extensively studied, but also scores particularly high in terms of genetic and linguistic congruence. "This might have given the impression that gene-language matches are the norm, but our study shows that this isn't the case," concludes Chiara Barbieri, who adds that it is important to include genetic and linguistic data from populations all over the world to understand language evolution.
The work is published in Proceedings of the National Academy of Sciences.
Written by Jan Bartek - AncientPages.com Staff Writer